Loading...
Interneurons migration in developing mammal brains
Abstract
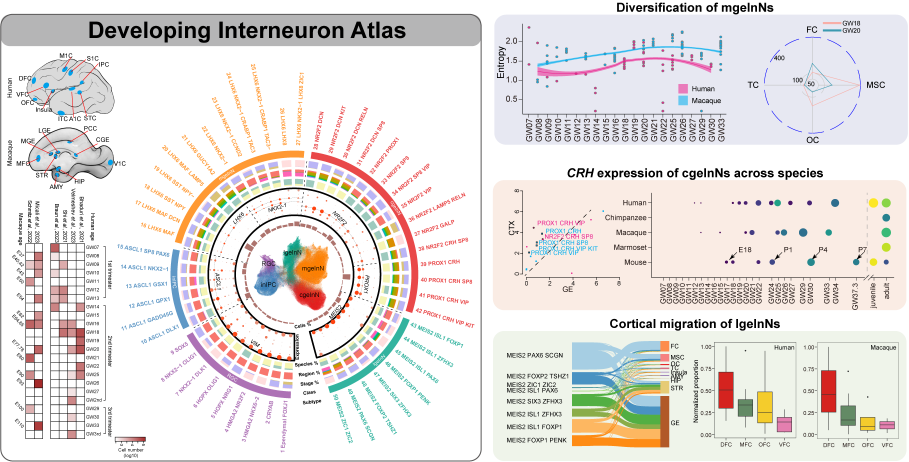
Cortical interneurons generated from ganglionic eminence via a long-distance journey of tangential migration display evident cellular and molecular differences across brain regions, which seeds the heterogeneous cortical circuitry in primates. However, whether such regional specifications in interneurons are intrinsically encoded or gained through interactions with the local milieu remains elusive. Here, we recruit 685,692 interneurons from cerebral cortex and subcortex including ganglionic eminence within the developing human and macaque species. Our integrative and comparative analyses reveal that less transcriptomic alteration is accompanied by interneuron migration within the ganglionic eminence subdivisions, in contrast to the dramatic changes observed in cortical tangential migration, which mostly characterize the transcriptomic specification for different destinations and for species divergence. Moreover, the in-depth survey of temporal regulation illustrates species differences in the developmental dynamics of cell types, e.g., the employment of CRH in primate interneurons during late-fetal stage distinguishes from their postnatal emergence in mice, and our entropy quantifications manifest the interneuron diversities gradually increase along the developmental ages in human and macaque cerebral cortices. Overall, our analyses depict the spatiotemporal features appended to cortical interneurons, providing a new proxy for understanding the relationship between cellular diversity and functional progression.
Citation
If you use the data in your research, please cite:
Feng, X*.,Gao, Y*., et al. Cortical arealization of interneurons defines shared and distinct molecular programs in developing human and macaque brains. Nature Communications (2025). https://doi.org/10.1038/s41467-025-56058-8
Contact
Mingfeng Li (mingfengli@hust.edu.cn); Huazhong University of Science and Technology
Loading...